RESEARCH INTERESTS
Our laboratory utilizes fluorescence methodologies to elucidate dynamic aspects of biomolecules. We are currently studying dynamin, a large (98kDa) GTPase which functions to “pinch-off” membrane vesicles in pathways such as receptor mediated endocytosis and synaptic vesicle recycling. We carry out both in vitro and in vivo studies on the self-association modes of dynamin as well as its interaction with membranes and other proteins such as endophilin and Arc/Arg3.1. We have also been studying dynamins with mutations that cause motor disorders, specifically Centronuclear Myopathy and Charcot-Marie-Tooth disease. These studies, both in vitro and in living cells, are aimed at understanding the molecular basis for dynamin’s involvement in these disorders. Recently, we began a project to study, both in vitro and in vivo, the protein Arc/Arg3.1, an immediate-early gene product which is a key regulator of synaptic plasticity, and is required for the formation of long-term spatial, taste and fear memories. We continue to develop applications of the phasor method, a visual approach to treatment of time-resolved fluorescence data, to in vitro systems such as intrinsic protein fluorescence and membrane systems.
TECHNIQUES:
Steady-state and time-resolved fluorescence
Phasors
Polarization/Anisotropy
Förster Resonance Energy Transfer
Two-Photon Fluorescence Microscopy; FCS, N&B, FLIM
Total Internal Reflection Fluorescence (TIRF) Microscopy
SELECTED FIGURES FROM RECENT PUBLICATIONS
Figure 6 from: Dimeric endophilin A2 stimulates assembly and GTPase activity of dynamin 2. Ross JA, Chen Y, Müller J, Barylko B, Wang L, Banks HB, Albanesi JP, Jameson DM. Biophys J. 2011 Feb 2;100(3):729-37.
Figure 6 from: Applications of phasors to in vitro time-resolved fluorescence measurements Stefl, M., James, N.G., Ross, J.A. and Jameson, D.M. (2011) Anal. Biochem. 410:62-69. 
Figure 6 from: Applications of phasor plots to in vitro protein studies. James NG, Ross JA, Stefl M, Jameson DM.Anal Biochem. 2011 Mar 1;410(1):70-6.
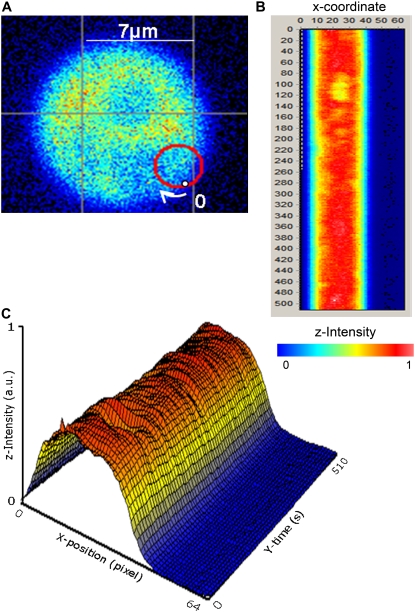
Figure 2 from: Oligomerization state of dynamin 2 in cell membranes using TIRF and number and brightness analysis. Ross JA, Digman MA, Wang L, Gratton E, Albanesi JP, Jameson DM. Biophys J. 2011 Feb 2;100(3):L15-7.
Figure 4 from: Fluorescence polarization/anisotropy in diagnostics and imaging. Jameson DM, Ross JA.Chem Rev. 2010 May 12;110(5):2685-708.
Figure 6 from: Characterization of Förster resonance energy transfer in a botulinum neurotoxin protease assay. Ross JA, Gilmore MA, Williams D, Aoki KR, Steward LE, Jameson DM.Anal Biochem. 2011 Jun 1;413(1):43-9.
Figure 1 from: Depolarization after resonance energy transfer (DARET): A sensitive fluorescence-based assay for botulinum neurotoxin protease activity. Gilmore MA, Williams D, Okawa Y, Holguin B, James NG, Ross JA, Roger Aoki K, Jameson DM, Steward LE.Anal Biochem. 2011 Jun 1;413(1):36-42.
Graphical abstract from: The proline/arginine-rich domain is a major determinant of dynamin self-activation. Barylko B, Wang L, Binns DD, Ross JA, Tassin TC, Collins KA, Jameson DM, Albanesi JP.Biochemistry. 2010 Dec 21;49(50):10592-4.
Figure 3A and B from: James, N.G., Ross, J.A., Mason, A.B. and Jameson, D.M. (2010) Protein Sci. 19:99-110. Excited-state lifetime studies of the three tryptophan residues in the N-lobe of human serum transferrin.
Figure 3a and b from: Fluorescence fluctuation spectroscopy: ushering in a new age of enlightenment for cellular dynamics. Jameson, D.M., Ross, J.A. and Albanesi, J.P. (2009) Biophysical Reviews 1: 105-118. 
 Figure 5 from: Martin, G.L., Ross, J.A., Minteer, S.D., Jameson, D.M. and Cooney, M.J. (2009) Carbo. Poly. 77: 695-702. Fluorescence Characterization of Chemical Microenvironments in Hydrophobically Modified Chitosan.
Figure 5 from: Martin, G.L., Ross, J.A., Minteer, S.D., Jameson, D.M. and Cooney, M.J. (2009) Carbo. Poly. 77: 695-702. Fluorescence Characterization of Chemical Microenvironments in Hydrophobically Modified Chitosan.
Figure 3 from: Yeast ribosomal stalk heterogeneity in vivo shown by two-photon FCS and molecular brightness analysis.García-Marcos, A., Sánchez, S.A., Parada, P., Eid, J., Jameson, D.M., Remacha, M., Gratton, E. and Ballesta, J.P.G. (2008) Biophysical J. 94:2884-2890.
 Figure 2 from: Arbildua, J.J., Brunet, J.E., Jameson, D.M., Lopez, M., Nova, E., Lagos, R. and Monasterio, O. (2006) Protein Sci. 15:410-9. Fluorescence resonance energy transfer and molecular modeling studies on 4′,6-diamidino-2-phenylindole (DAPI) complexes with tubulin.
Figure 2 from: Arbildua, J.J., Brunet, J.E., Jameson, D.M., Lopez, M., Nova, E., Lagos, R. and Monasterio, O. (2006) Protein Sci. 15:410-9. Fluorescence resonance energy transfer and molecular modeling studies on 4′,6-diamidino-2-phenylindole (DAPI) complexes with tubulin. Figure 2. Changes in anisotropy (A) and DAPI fraction bound (B) during DAPI titration with tubulin, tubulin S, and the complex DAPI–tubulin S with the C-terminal peptides. Titration of 0.5 μM DAPI with tubulin (•) and tubulin S (○). The buffer used was 10 mM Tris-HCl (pH 7.5). The inset shows the titration of 0.5 μM DAPI plus 1.5 μM tubulin S with increasing concentrations of C-terminal peptides (▴). Excitation wavelength 350 nm; emission wavelength 455 nm; excitation and emission bandwidths were 2.5 and 12 nm, respectively.
Figure 2. Changes in anisotropy (A) and DAPI fraction bound (B) during DAPI titration with tubulin, tubulin S, and the complex DAPI–tubulin S with the C-terminal peptides. Titration of 0.5 μM DAPI with tubulin (•) and tubulin S (○). The buffer used was 10 mM Tris-HCl (pH 7.5). The inset shows the titration of 0.5 μM DAPI plus 1.5 μM tubulin S with increasing concentrations of C-terminal peptides (▴). Excitation wavelength 350 nm; emission wavelength 455 nm; excitation and emission bandwidths were 2.5 and 12 nm, respectively.